STRING
Getting Start 入门
Starting Point 开始
To specify your desired starting point of the analysis you have to use the input form at the STRING start page.
要指定所需的分析起点,您必须使用 STRING 起始页上的输入表单。
- Protein by name 蛋白质名称
- Protein by sequence 按序列划分的蛋白质
- Multiple proteins 多种蛋白质
- Multiple sequences 多个序列
- Organisms 生物体
- Protein families (COGs) 蛋白质家族 (COG)
- Examples 例子
- Random entry 随机输入
You can search STRING by single protein name, multiple names or by amino acid sequence (in any format) There are also example inputs and a random input generator which will randomly select a protein with at least 4 predicted links at medium confidence or better. There is a organism entry to see if your species of interest is available. There is the possibility to search by protein family rather than a protein in a single organism, by searching the COGs (clusters of orthologous groups)
您可以按单个蛋白质名称、多个名称或氨基酸序列(任何格式)搜索 STRING,还有一些示例输入和随机输入生成器,它将以中等置信度或更高随机选择具有至少 4 个预测链接的蛋白质。有一个生物条目,可以查看您感兴趣的物种是否可用。通过搜索COG(直系同源组簇),可以按蛋白质家族而不是单个生物体中的蛋白质进行搜索
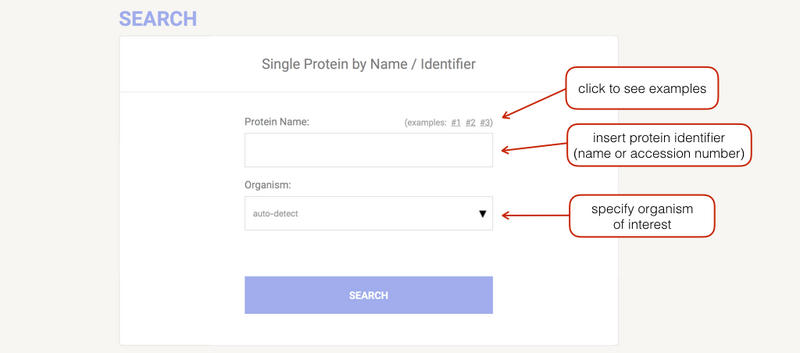
Commonly, you enter your protein of interest by supplying its name or identifier. The organism can be selected by clicking on the arrow or directly typing the name inside the relative input field (an autocompletion mechanism will appear to help you). General names that group more than one organism (e.g. "Mammals", "Chordata") can also be used.
通常,您可以通过提供其名称或标识符来输入目标蛋白质。可以通过单击箭头或直接在相关输入字段中输入名称来选择生物体(将出现自动完成机制来帮助您)。也可以使用将一种以上生物分组的通用名称(例如“哺乳动物”、“脊索动物”)。
Network 网络
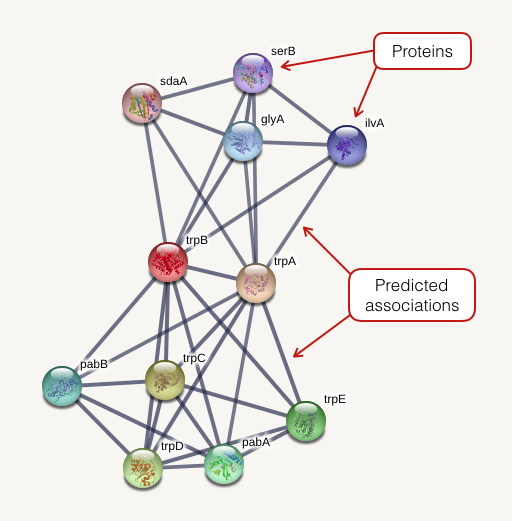
The network view summarizes the network of predicted associations for a particular group of proteins. The network nodes are proteins. The edges represent the predicted functional associations. The edges are draw according to the view settings. In evidence mode, an edge may be drawn with up to 7 differently colored lines - these lines represent the existence of the seven types of evidence used in predicting the associations.
网络视图汇总了特定蛋白质组的预测关联网络。网络节点是蛋白质。边表示预测的功能关联。边是根据视图设置绘制的。在证据模式下,最多可以用 7 条不同颜色的线绘制一条边 - 这些线代表用于预测关联的七种证据的存在。
- Red line - indicates the presence of fusion evidence 红线 - 表示存在融合证据
- Green line - neighborhood evidence 绿线 - 邻域证据
- Blue line - cooccurrence evidence 蓝线 - 共现证据
- Purple line - experimental evidence 紫线——实验证据
- Yellow line - textmining evidence 黄线 - 文本挖掘证据
- Light blue line - database evidence 浅蓝线 - 数据库证据
- Black line - coexpression evidence. 黑线 - 共表达证据。
In confidence mode the thickness of the line indicate the degree of confidence prediction of the interaction. Action mode show additional information about the prediction, such as, binding, activation, etc.
在置信模式下,线的粗细表示交互作用预测的置信度。操作模式显示有关预测的其他信息,例如绑定、激活等。
Clicking on a node gives several details about the protein. Clicking on an edge displays a detailed evidence breakdown.
单击节点可提供有关蛋白质的几个详细信息。单击边缘会显示详细的证据细分。
A note on the network drawing algorithm
关于网络绘制算法的说明
STRING uses a spring model to generate the network images. Nodes are modeled as masses and edges as springs; the final position of the nodes in the image is computed by minimizing the 'energy' of the system. We give high confidence edges a higher 'spring strength' so that they will reach an optimal position before lower confidence edges. The user also can optionally reduce the 'natural length' of a high confidence edge - this forces them closer together and sometimes results in a clearer picture of high confidence interactions. We set the high confidence edge length to 80% of the normal length by default.
STRING 使用弹簧模型来生成网络图像。节点被建模为质量,边被建模为弹簧;图像中节点的最终位置是通过最小化系统的“能量”来计算的。我们赋予高置信度边缘更高的“弹簧强度”,以便它们在低置信度边缘之前达到最佳位置。用户还可以选择减少高置信度边缘的“自然长度”——这迫使它们靠得更近,有时会导致更清晰的高置信度交互画面。默认情况下,我们将高置信度边长度设置为正常长度的 80%。
This modeling has some important consequences that the user should be aware of. Firstly, the physical distances between two nodes along an edge in a graph has no meaning; indeed, an attempt to set the edge length based on score would probably result in an unsolvable set of equations! We try to ensure high confidence links are drawn close together through the setting of the modeling parameters described above. Secondly, although the algorithm is deterministic - the same input will produce the same output - the addition of, say, new nodes to the network can result in node locations in the new image completely changing. Finally, although the input node is the 'center' of the network in an abstract sense, it may not be located centrally in the network image.
这种建模具有用户应注意的一些重要后果。首先,图中沿边的两个节点之间的物理距离没有意义;事实上,尝试根据分数设置边长可能会导致一组无法解决的方程!我们试图通过设置上述建模参数来确保高置信度链接紧密地联系在一起。其次,尽管该算法是确定性的——相同的输入将产生相同的输出——但向网络中添加新节点可能会导致新图像中的节点位置完全改变。最后,尽管输入节点在抽象意义上是网络的“中心”,但它可能并不位于网络图像的中心位置。
The navigation take you to different aspects of the data, allowing you change parameters and to see the different types of evidence that supports the predicted associations.
导航将您带到数据的不同方面,允许您更改参数并查看支持预测关联的不同类型的证据。
Legend 图例
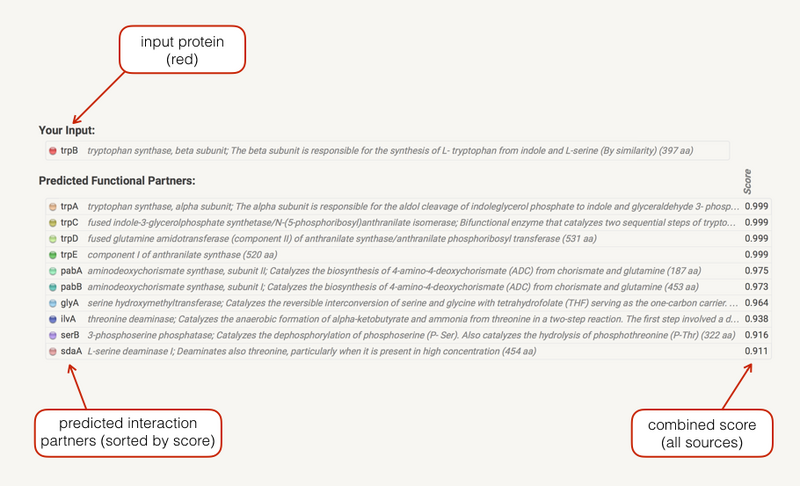
In the Legend section a list your input(s) is shown. Predicted associations are shown immediately in a list below your input, sorted by score. If input gene is a fusion of two functions, both will be shown. Clicking on the score bullets gives you a breakdown of the individual prediction method scores. Clicking on a gene name gives you the protein sequence as well as a list of similar proteins in STRING. Initially, only predictions with medium (or better) confidence, limited to the top 10 interactors will be shown. These parameters can be changed in the data settings.
在“图例”部分中,将显示您的输入列表。预测的关联会立即显示在输入下方的列表中,并按分数排序。如果输入基因是两种功能的融合,则将显示两者。单击分数项目符号可显示各个预测方法分数的细分。单击基因名称可为您提供蛋白质序列以及 STRING 中类似蛋白质的列表。最初,仅显示具有中等(或更高)置信度的预测,仅限于前 10 个交互者。这些参数可以在数据设置中更改。
Data Settings 数据设置
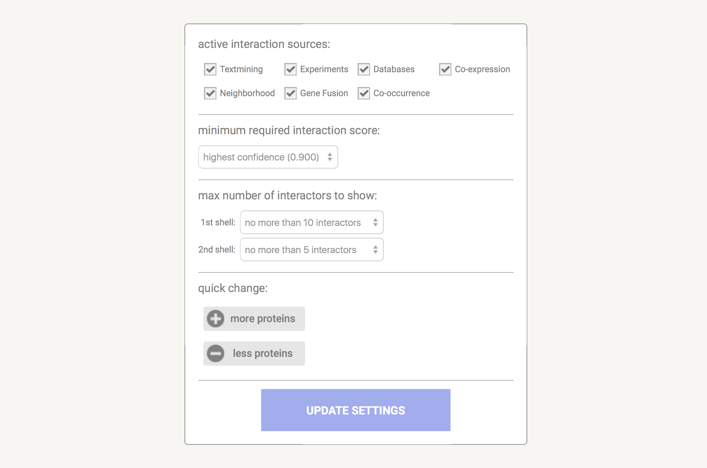
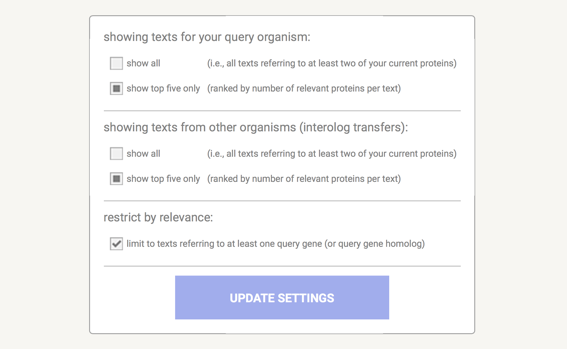
In the data settings you can change the parameters that influence the output. Note that parameters are only changed when you press the 'Update Settings' button.
在数据设置中,您可以更改影响输出的参数。请注意,仅当您按下“更新设置”按钮时,参数才会更改。
Under the active interaction sources you can select which type of evidence will contribute to the prediction of the score.
在主动交互源下,您可以选择哪种类型的证据将有助于分数的预测。
The minimum required interaction score puts a threshold on the confidence score, such that only interaction above this score are included in the predicted network. Lower score mean more interaction, but also more false positives. The confidence score is the approximate probability that a predicted link exists between two enzymes in the same metabolic map in the KEGG database. Confidence limits are as follows:
所需的最低交互分数在置信度分数上设置了一个阈值,因此只有高于此分数的交互才会包含在预测网络中。分数越低意味着交互越多,但误报也越多。置信度分数是 KEGG 数据库中同一代谢图谱中两种酶之间存在预测链接的近似概率。置信限如下:
- low confidence - 0.15 (or better), 低置信度 - 0.15(或更好),
- medium confidence - 0.4, 中等置信度 - 0.4,
- high confidence - 0.7, 高置信度 - 0.7,
- highest confidence - 0.9. 最高置信度 - 0.9。
You can choose the max number of interactions to show. This is an option to limit the number of interactions to your input. The default setting is to limit the output to the 10 best-scoring hits.
您可以选择要显示的最大互动次数。此选项用于限制与输入的交互次数。默认设置是将输出限制为 10 个得分最高的命中。
There is a options the set how many interactions are shown that directly connect with your input by setting the 1st shell and how many indirect interaction that connect to a protein in the first shell by setting the 2nd shell. Please note that this can result in fairly large networks that may take a while to compute and download.
有一个选项:通过设置第一个壳来设置与你的输入直接连接的相互作用,以及通过设置第二个壳来设置与第一个壳中的蛋白质连接的间接相互作用。请注意,这可能会导致相当大的网络,可能需要一段时间来计算和下载。
Note that you can click on any node, and the subsequent page offers a link to use that node as the input - effectively placing it in the center of the image. Repeated use of this mechanism allows you to explore large regions of the network.
请注意,您可以单击任何节点,随后的页面会提供一个链接,用于将该节点用作输入 - 有效地将其放置在图像的中心。重复使用此机制可以探索网络的大区域。
You have the convenient option the change the parameter setting by pressing either of the buttons und quick change.
您可以通过按任一按钮和快速更改来更改参数设置。
View Settings 视图设置
The dialogue box shown above is the one for the Network View. Network specific parameters are: 'edge scaling factor' - this reduces the length of high-scoring edges so that the images will be drawn more compact, and low scoring hits will be spread out further. Lower values mean more compact images, higher values will cause more spread.
上面显示的对话框是网络视图的对话框。特定于网络的参数是:“边缘缩放因子” - 这减少了高分边缘的长度,以便绘制更紧凑的图像,并且低分命中将进一步分散。较低的值意味着更紧凑的图像,较高的值将导致更多的扩散。
Here you can select the meaning of network edges of the displayed network. You can choose between:
在这里,您可以选择所显示网络的网络边缘的含义。您可以选择:
- evidence - multiple lines where the color indicates the type of interaction evidence 证据 - 多行,其中颜色表示交互证据的类型
- confidence - line thickness indicates the strength of data support 置信度 - 线粗表示数据支持的强度
- molecular action - line shape indicates the predicted mode of action 分子作用 - 线形表示预测的作用模式
The network display mode gives you the option to change the format of the displayed network. The options are:
网络显示模式为您提供了更改显示网络格式的选项。选项有:
- static png - image of network is a simple bitmap image that is not interactive 静态 PNG - 网络图像是非交互式的简单位图图像
- interactive svg - network is a scalable vector graphic [SVG] and provides interactivity 交互式 SVG - 网络是可缩放的矢量图形 [SVG],提供交互性
- interactive flash - network is displayed in a Flash-applet, which allows for functionality not (yet) implemented in svg mode (e.g. clustering) 交互式 Flash - 网络显示在 Flash 小程序中,它允许(尚未)在 svg 模式下实现的功能(例如集群)
Finally, you can disable structure previews inside network bubbles by checking the check-box protein structure information.
最后,您可以通过选中复选框蛋白质结构信息来禁用网络气泡内的结构预览。
Tables / Exports 表格/导出
In this section you can export your current network to the following formats:
在本节中,您可以将当前网络导出为以下格式:
- bitmap image - image of the network in the PNG (portable network graphic) file format. 位图图像 - PNG(可移植网络图形)文件格式的网络图像。
- high-resolution bitmap - image in PNG format, at resolution 400 dpi 高分辨率位图 - PNG 格式的图像,分辨率为 400 dpi
- vector graphic: - image in SVG (scalable vector graphic) format that can be opened and edited in Illustrator, CorelDraw, Dia, etc. 矢量图形: - SVG(可缩放矢量图形)格式的图像,可以在 Illustrator、CorelDraw、Dia 等中打开和编辑。
- simple tabular text output - data for the interaction network as tab separated values (TSV format). File can be opened in Excel. This data is also show further down on Table/Export page. 简单的表格文本输出 - 以制表符分隔值(TSV 格式)的形式进行交互网络的数据。文件可以在Excel中打开。此数据也会显示在“表/导出”页面上的下方。
- XML summary - interaction data in a structured XML data format, according to the 'PSI-MI' data standard XML 摘要 - 根据“PSI-MI”数据标准,结构化 XML 数据格式的交互数据
- network coordinates - a flat-file format describing the coordinates and colors of nodes in the network 网络坐标 - 描述网络中节点坐标和颜色的平面文件格式
- protein sequences - MFA: multi-fasta format, containing the amino acid sequences in the network 蛋白质序列 - MFA:multi-fasta 格式,包含网络中的氨基酸序列
- protein annotations - a tab-delimited file describing the names, domains and annotated functions of the network proteins 蛋白质注释 - 一个制表符分隔的文件,描述网络蛋白质的名称、结构域和注释功能
Evidence 证据
Conserved Neighborhood 保守邻域
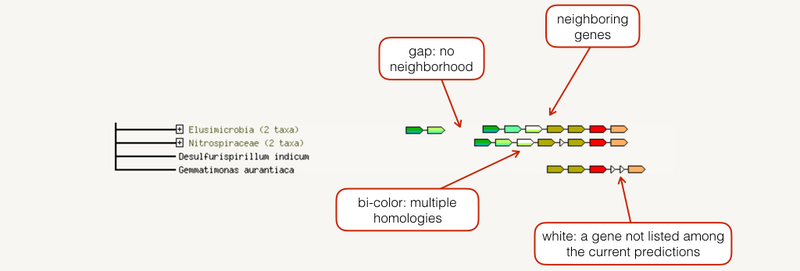
The neighborhood view shows runs of genes that occur repeatedly in close neighborhood in (prokaryotic) genomes. Genes located together in a run are linked with a black line (maximum allowed intergenic distance is 300 base pairs). Note that if there are multiple runs for a given species, these are separated by white space. If there are other genes in the run that are below the current score threshold, they are drawn as small white triangles. Gene fusion occurrences are also drawn, but only if they are present in a run (see also the Fusion section below for more details).
邻域视图显示了在(原核)基因组中在近邻域中重复出现的基因运行。在运行中位于一起的基因用黑线连接(最大允许的基因间距离为 300 个碱基对)。请注意,如果给定物种有多个游程,则这些游程由空格分隔。如果运行中还有其他低于当前分数阈值的基因,则将它们绘制为白色小三角形。还会绘制基因融合事件,但前提是它们在运行中存在(有关详细信息,另请参阅下面的融合部分)。
Co-occurrence 共现
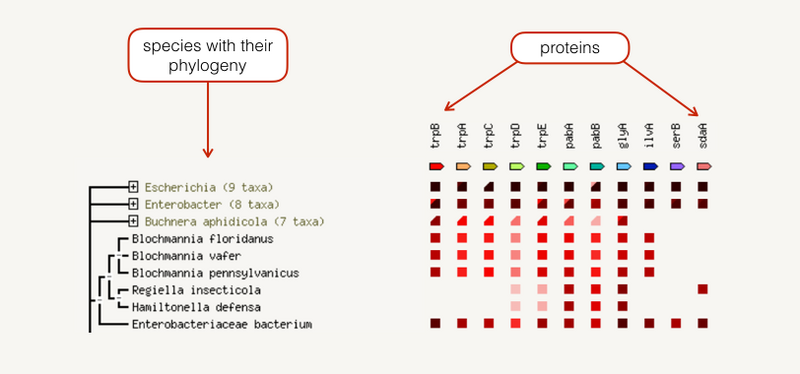
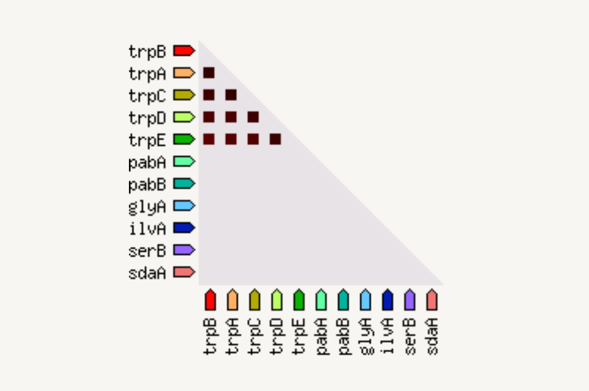
The occurrence view shows the presence or absence of linked proteins across species. Proteins are listed across the top of the page and a phylogenetic tree with species names is listed down the left hand side. In the subsequent grid, the presence of the protein in a species is marked with a red square and absence with a white space. The intensity of the color of the red square reflect the amount of conservation of the homologous protein in the specie.
发生视图显示跨物种是否存在连接蛋白。蛋白质列在页面顶部,带有物种名称的系统发育树列在左侧。在随后的网格中,物种中蛋白质的存在用红色方块标记,而不存在则用空白标记。红色方块颜色的强度反映了物种中同源蛋白质的保守程度。
Fusion 融合
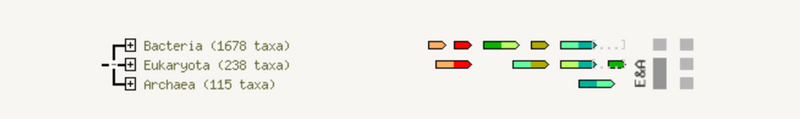
The fusion view shows the individual gene fusion events per species. The species in which fusion occurs are listed to the left. Genes are colored according to the table at the bottom of the page. White genes are those which are fused but not directly linked to the input at the selected confidence level. Hovering above a region in a gene gives the gene name; clicking on a gene gives more detailed information.
融合视图显示每个物种的单个基因融合事件。发生融合的物种列在左侧。基因根据页面底部的表格着色。白色基因是那些融合但未直接链接到所选置信水平的输入的基因。将鼠标悬停在基因中的某个区域上方可获得基因名称;点击基因可以提供更详细的信息。
Co-expression 共表达
The coexpression view shows the genes that are co-expressed in the same or in other species (transferred by homology). Co-expression is shown by a red square: more intense color of the square represent a higher association score of the expression data.
共表达视图显示在同一物种或其他物种中共表达的基因(通过同源性转移)。共表达用红色方块表示:方块颜色越深表示表达数据的关联分数越高。
Experiments 实验
The experiments view shows a list of significant protein interaction datasets, gathered from other protein-protein interaction databases. The name of the database is present in the grey header of the table: you can get more information on the group, clicking on the "info" link. Below the header, the organism is reported together with the proteins of the network that are present in this group.
实验视图显示了从其他蛋白质-蛋白质相互作用数据库收集的重要蛋白质相互作用数据集的列表。数据库的名称显示在表格的灰色标题中:您可以单击“信息”链接以获取有关该组的更多信息。在标题下方,生物体与该组中存在的网络蛋白质一起报告。
Databases 数据库
This view shows a list of significant protein interaction groups, gathered from curated databases. You can get more information on the group, clicking on the "info" link on the grey rows. Clicking the bubbles next to their respective gene names give information of the individual proteins.
此视图显示了从精选数据库中收集的重要蛋白质相互作用组的列表。您可以获得有关该组的更多信息,单击灰色行上的“信息”链接。单击它们各自基因名称旁边的气泡可提供单个蛋白质的信息。
Text mining 文本挖掘
The text mining view shows a list of significant protein interaction groups, extracted from the abstracts of scientific literature. The title and the abstract of the publication are displayed together with a link to the publication.
文本挖掘视图显示了从科学文献摘要中提取的重要蛋白质相互作用基团的列表。出版物的标题和摘要与出版物的链接一起显示。
Analysis 分析
The analysis section give some brief statistics of the inferred network, such as the number of nodes and edges. The average node degree is a number of how many interactions (at the score threshold) that a protein have on the average in the network. The clustering coefficient is a measure of how connected the nodes in the network are.
分析部分给出了推断网络的一些简要统计数据,例如节点数和边数。平均节点度是蛋白质在网络中平均有多少相互作用(在分数阈值下)的数字。聚类系数是衡量网络中节点连接程度的指标。
Highly connected networks have high values.
高度互联的网络具有很高的价值。
The expected number of edges gives how many edges is to be expected if the nodes were to be selected at random. A small PPI enrichment p-value indicate that the nodes are not random and that the observed number of edges is significant. Note that is some cases enrichment is to be expected and that there numbers have to be interpreted with some caution.
预期的边数表示,如果随机选择节点,预期的边数。较小的 PPI 富集 p 值表明节点不是随机的,并且观察到的边数很大。请注意,在某些情况下,浓缩是可以预料的,并且必须谨慎地解释数字。
There is also an enrichment analysis for Gene Ontologies, pathways and domains. Basically, this shows term that are more enriched in the set of proteins in the network than the background.
还有针对基因本体、通路和结构域的富集分析。基本上,这显示了在网络中蛋白质集中比背景更丰富的术语。