Omicscloud配置
mariadb数据库
安装:
sudo apt update
sudo apt install mariadb-server mariadb-client -y
安装后启动:
sudo systemctl start mariadb
sudo systemctl status mariadb
数据库导入
记得改cd文件夹名称或数据库名称。
cd /stor2/db_bk_20260324 && \
sudo mariadb -e "CREATE DATABASE IF NOT EXISTS fasta_db CHARACTER SET utf8mb4 COLLATE utf8mb4_general_ci; CREATE DATABASE IF NOT EXISTS idmapping CHARACTER SET utf8mb4 COLLATE utf8mb4_general_ci; CREATE DATABASE IF NOT EXISTS kegg_mtable CHARACTER SET utf8mb4 COLLATE utf8mb4_general_ci; CREATE DATABASE IF NOT EXISTS omicscloud CHARACTER SET utf8mb4 COLLATE utf8mb4_general_ci; CREATE DATABASE IF NOT EXISTS speciesdb_info CHARACTER SET utf8mb4 COLLATE utf8mb4_general_ci; CREATE DATABASE IF NOT EXISTS userdb CHARACTER SET utf8mb4 COLLATE utf8mb4_general_ci;" && \
sudo mariadb fasta_db < fasta_db_dump_20260324.sql && \
sudo mariadb idmapping < idmapping_dump_20260324.sql && \
sudo mariadb kegg_mtable < kegg_mtable_dump_20260324.sql && \
sudo mariadb omicscloud_db < omicscloud_db_dump_20260324.sql && \
sudo mariadb speciesdb_info < speciesdb_info_dump_20260324.sql && \
sudo mariadb userdb < userdb_dump_20260324.sql
导完后检查一下:
sudo mariadb -e "SHOW DATABASES;"
复制KEGG.7z到挂载文件夹
mkdir -p ~/mnt/database/KEGG
cp /stor2/db_bk_20260324/KEGG.7z ~/mnt/database/KEGG
设置root密码
sudo mariadb
ALTER USER 'root'@'localhost' IDENTIFIED BY '你的新密码';
GRANT ALL PRIVILEGES ON my_data.* TO 'omicscloud'@'localhost';
FLUSH PRIVILEGES;
exit;
DBeaver
下载DBeaver-ce并登录omicscloud账户,可查看和修改表格
LDAP
docker安装
sudo apt remove -y docker.io docker-compose docker-compose-v2 podman-docker containerd runc
sudo apt update
sudo apt install -y ca-certificates curl
sudo install -m 0755 -d /etc/apt/keyrings
sudo curl -fsSL https://download.docker.com/linux/ubuntu/gpg -o /etc/apt/keyrings/docker.asc
sudo chmod a+r /etc/apt/keyrings/docker.asc
echo "deb [arch=$(dpkg --print-architecture) signed-by=/etc/apt/keyrings/docker.asc] https://download.docker.com/linux/ubuntu $(. /etc/os-release && echo ${UBUNTU_CODENAME}) stable" | sudo tee /etc/apt/sources.list.d/docker.list > /dev/null
sudo apt update
sudo apt install -y docker-ce docker-ce-cli containerd.io docker-buildx-plugin docker-compose-plugin
sudo systemctl enable --now docker
sudo systemctl status docker
sudo docker run hello-world
docker compose version
如果docker网络有问题:
sudo tee /etc/resolv.conf > /dev/null <<'EOF'
nameserver 223.5.5.5
nameserver 119.29.29.29
nameserver 8.8.8.8
EOF
sudo systemctl restart docker
创建docker-compose.yml
services:
#ldap服务
openldap:
image: osixia/openldap
container_name: openldap-server
hostname: ldap-server
restart: always
networks:
- ldap
ports:
- '389:389'
- '636:636'
volumes:
- /opt/openldap/ldap:/var/lib/ldap
- /opt/openldap/slapd.d:/etc/ldap/slapd.d
- /opt/openldap/certs:/container/service/slapd/assets/certs
environment:
- LDAP_ORGANISATION=omicsolution #组织名称,需要改
- LDAP_DOMAIN=omicsolution.com #域名,需要改
- LDAP_ADMIN_USERNAME=admin
- LDAP_ADMIN_PASSWORD=123456
#- LDAP_USERS=user01,user02
#- LDAP_PASSWORDS=password1,password2
#页面管理
phpldapadmin:
image: osixia/phpldapadmin
container_name: openldap-admin
hostname: ldap-admin
restart: always
privileged: true #授予真实root权限
networks:
- ldap
ports:
- '50080:80'
#- '443:443' #PHPLDAPADMIN_HTTPS为true有效
environment:
- PHPLDAPADMIN_HTTPS=false
- PHPLDAPADMIN_LDAP_HOSTS=ldap-server #指向openldap的hostname
depends_on:
- openldap
networks:
ldap:
#driver: bridge
装好后,进入 docker-compose.yml 所在目录,启动:
sudo mkdir -p /opt/openldap/ldap
sudo mkdir -p /opt/openldap/slapd.d
sudo mkdir -p /opt/openldap/certs
sudo docker compose -p ldap -f /home/lyx/文档/docker-compose.yml up -d
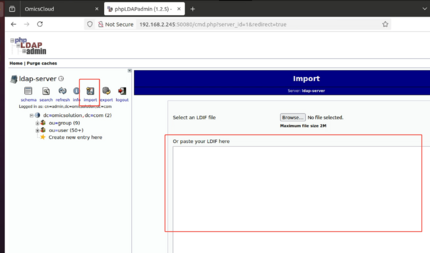
LDAP导入
把 ldif-export.txt 导进去:
phpldapadmin登录的用户名是cn=admin,dc=omicsolution,dc=com
密码在yml文件里
勾上Don't stop on errors
环境需要的软件
R和shiny
sudo tee /etc/apt/sources.list.d/ubuntu.sources > /dev/null <<'EOF'
Types: deb
URIs: http://archive.ubuntu.com/ubuntu/
Suites: noble noble-updates noble-backports
Components: main restricted universe multiverse
Signed-By: /usr/share/keyrings/ubuntu-archive-keyring.gpg
Types: deb
URIs: http://security.ubuntu.com/ubuntu/
Suites: noble-security
Components: main restricted universe multiverse
Signed-By: /usr/share/keyrings/ubuntu-archive-keyring.gpg
EOF
sudo apt update
sudo apt --fix-broken install
sudo apt full-upgrade -y
sudo apt install -y r-base r-base-dev
R --version
sudo R -e "install.packages('shiny', repos='https://cran.rstudio.com/')"
sudo apt install -y gdebi-core
cd /tmp
wget https://download3.rstudio.org/ubuntu-20.04/x86_64/shiny-server-1.5.23.1030-amd64.deb
sudo gdebi -n shiny-server-1.5.23.1030-amd64.deb
sudo systemctl enable --now shiny-server
sudo systemctl status shiny-server
安装R包
ubuntu依赖:
sudo apt update
sudo apt-get install libcurl4-openssl-dev libssl-dev libxml2-dev default-jdk
sudo apt install -y \
build-essential gfortran cmake make \
libcurl4-openssl-dev libssl-dev libxml2-dev \
libcairo2-dev libxt-dev pkg-config \
libfontconfig1-dev libfreetype6-dev \
libharfbuzz-dev libfribidi-dev \
libjpeg-dev libpng-dev libtiff5-dev libglib2.0-dev \
libglu1-mesa-dev freeglut3-dev mesa-common-dev \
default-jdk r-base-dev
sudo apt install -y \
build-essential gfortran pkg-config \
libgl1-mesa-dev libglu1-mesa-dev freeglut3-dev mesa-common-dev \
libx11-dev libxt-dev libxmu-dev libxi-dev \
default-jdk openjdk-17-jdk-headless
sudo apt install -y libtirpc-dev default-jdk openjdk-17-jdk-headless
sudo R CMD javareconf
R包: 下载安装r-studio
# 安装所有包(智能检测源:优先 CRAN,失败后尝试 Bioconductor)
# 运行前请确保已安装 BiocManager
# 首先安装 BiocManager(如果未安装)
if (!requireNamespace("BiocManager", quietly = TRUE)) {
install.packages("BiocManager")
}
# 定义所有包列表(合并所有包,去重)
all_packages <- unique(c(
# 原 CRAN 包
"abind", "ade4", "ape", "aplot", "AsioHeaders", "askpass", "async",
"backports", "base64url", "BH", "bit", "bit64", "bitops", "blob",
"brew", "brio", "broom", "Cairo", "calibrate", "callr", "car",
"carData", "caTools", "cellranger", "chromote", "chron", "clipr",
"clock", "colorspace", "colourpicker", "conflicted", "cookies",
"corpcor", "corrplot", "countrycode", "cowplot", "cpp11", "credentials",
"crosstalk", "curl", "data.table", "DBI", "dbplyr", "dendextend",
"Deriv", "desc", "devtools", "diffobj", "doBy", "doParallel", "doRNG",
"downlit", "downloader", "dplyr", "dragulaR", "DT", "dtplyr",
"echarts4r", "ellipse", "ellipsis", "emmeans", "estimability",
"evaluate", "factoextra", "FactoMineR", "fansi", "farver", "flashClust",
"forcats", "foreach", "formatR", "Formula", "fresh", "futile.logger",
"futile.options", "future", "gargle", "gdata", "generics", "gert",
"getopt", "ggdendro", "ggfun", "ggnewscale", "ggplot2", "ggplotify",
"ggpubr", "ggrepel", "ggsci", "ggseqlogo", "ggsignif", "ggstar", "gh",
"gitcreds", "globals", "googledrive", "googlesheets4", "GOplot",
"gplots", "gridBase", "gridExtra", "gridGraphics", "gsignal", "gsubfn",
"gtable", "gtools", "hash", "haven", "highr", "hms", "htmlwidgets",
"httr", "httr2", "ids", "igraph", "ini", "isoband", "iterators",
"itertools", "KEGGgraph", "knitr", "labeling", "lambda.r", "lazyeval",
"leaps", "listenv", "litedown", "lme4", "lubridate", "markdown",
"Matrix", "MatrixModels", "matrixStats", "mclust", "microbenchmark",
"miniUI", "minqa", "missForest", "mixOmics", "modelr", "multcompView",
"mvtnorm", "nloptr", "numDeriv", "ontologyIndex", "openssl", "openxlsx",
"optparse", "parallelly", "patchwork", "pathview", "pbkrtest",
"pdftools", "Peptides", "pheatmap", "pillar", "pixmap", "pkgbuild",
"pkgconfig", "pkgdown", "pkgload", "PKI", "plogr", "plotly", "plotrix",
"plyr", "png", "polynom", "pracma", "praise", "prettyunits",
"processx", "profvis", "progress", "proto", "ps", "purrr", "qpdf",
"qqman", "quantreg", "randomForest", "rARPACK", "rbibutils",
"rChoiceDialogs", "rcmdcheck", "RColorBrewer", "RcppArmadillo",
"RcppEigen", "RCurl", "Rdpack", "reactR", "reactwidgets", "readr",
"readxl", "reformulas", "relayer", "rematch", "rematch2", "remotes",
"reprex", "reshape", "reshape2", "rgl", "rhandsontable", "rJava",
"rjson", "RJSONIO", "rlist", "RMariaDB", "rmarkdown", "rngtools",
"ropls", "roxygen2", "RPostgreSQL", "rprojroot", "RSpectra", "RSQLite",
"rstatix", "rstudioapi", "rversions", "rvest", "scales",
"scatterplot3d", "segmented", "selectr", "seqinr", "sessioninfo",
"sfsmisc", "shiny.i18n", "shinyalert", "shinyBS", "shinybusy",
"shinycssloaders", "shinydashboard", "shinydashboardPlus", "shinyFiles",
"shinyjs", "shinysky", "shinythemes", "shinyWidgets", "simsalapar",
"snow", "sp", "SparseM", "sqldf", "statmod", "stringi", "stringr",
"svDialogs", "svGUI", "sys", "systemfonts", "testthat", "textshaping",
"tibble", "tidyr", "tidyselect", "tidytree", "tidyverse", "timechange",
"tinytex", "tzdb", "urlchecker", "usethis", "utf8", "uuid", "V8",
"vctrs", "VennDiagram", "viridis", "viridisLite", "visNetwork", "vroom",
"waiter", "waldo", "webshot2", "websocket", "wesanderson", "whisker",
"xfun", "XML", "xml2", "xopen", "yaml", "yulab.utils", "zip",
"base64enc", "bslib", "cachem", "cli", "commonmark", "crayon", "digest",
"fastmap", "fontawesome", "fs", "glue", "htmltools", "httpuv",
"jquerylib", "jsonlite", "later", "lifecycle", "magrittr", "memoise",
"mime", "promises", "R6", "rappdirs", "Rcpp", "rlang", "sass", "shiny",
"sourcetools", "withr", "xtable",
# 原 Bioconductor 包
"affy", "affyio", "AnnotationDbi", "Biobase", "BiocBaseUtils",
"BiocGenerics", "BiocManager", "BiocParallel", "BiocVersion",
"Biostrings", "DelayedArray", "GenomeInfoDb", "GenomeInfoDbData",
"GenomicRanges", "graph", "impute", "IRanges", "KEGGREST", "limma",
"MatrixGenerics", "MultiAssayExperiment", "MultiDataSet", "org.Hs.eg.db",
"preprocessCore", "qvalue", "Rgraphviz", "S4Arrays", "S4Vectors",
"SparseArray", "STRINGdb", "SummarizedExperiment", "UCSC.utils", "vsn",
"XVector"
))
# 检查包是否已安装
is_installed <- function(pkg) {
requireNamespace(pkg, quietly = TRUE)
}
# 从 CRAN 安装
install_from_cran <- function(pkg) {
install.packages(pkg, dependencies = FALSE, quiet = FALSE)
}
# 从 Bioconductor 安装
install_from_bioc <- function(pkg) {
BiocManager::install(pkg, update = FALSE, ask = FALSE, quiet = FALSE)
}
# 尝试从指定源安装,如果失败则返回 FALSE
try_install <- function(pkg, install_func, source_name) {
tryCatch({
cat(" 尝试从", source_name, "安装...\n")
install_func(pkg)
cat(" ✓ 从", source_name, "安装成功\n")
return(TRUE)
}, error = function(e) {
cat(" ✗ 从", source_name, "安装失败:", e$message, "\n")
return(FALSE)
})
}
# 智能安装函数:先尝试 CRAN,失败则尝试 Bioconductor
smart_install <- function(pkg) {
# 跳过已安装的包
if (is_installed(pkg)) {
cat("⊙ 已跳过:", pkg, "(已安装)\n")
return(TRUE)
}
cat("\n正在安装:", pkg, "\n")
# 先尝试从 CRAN 安装
if (try_install(pkg, install_from_cran, "CRAN")) {
return(TRUE)
}
# CRAN 失败,尝试从 Bioconductor 安装
cat(" CRAN 安装失败,尝试从 Bioconductor 安装...\n")
if (try_install(pkg, install_from_bioc, "Bioconductor")) {
return(TRUE)
}
# 都失败
cat(" ✗✗✗ 安装失败:", pkg, "(CRAN 和 Bioconductor 均失败)\n")
return(FALSE)
}
# 执行安装
cat("========================================\n")
cat("开始智能安装所有包...\n")
cat("安装策略: 优先 CRAN → 失败则尝试 Bioconductor\n")
cat("========================================\n\n")
# 记录安装结果
success_count <- 0
fail_count <- 0
failed_packages <- c()
for (pkg in all_packages) {
result <- smart_install(pkg)
if (result) {
success_count <- success_count + 1
} else {
fail_count <- fail_count + 1
failed_packages <- c(failed_packages, pkg)
}
}
# 输出统计信息
cat("\n\n========================================\n")
cat("安装完成!统计信息:\n")
cat("========================================\n")
cat("总包数:", length(all_packages), "\n")
cat("成功安装/已存在:", success_count, "\n")
cat("安装失败:", fail_count, "\n")
if (fail_count > 0) {
cat("\n失败的包列表:\n")
for (pkg in failed_packages) {
cat(" -", pkg, "\n")
}
cat("\n建议:\n")
cat("1. 检查网络连接\n")
cat("2. 检查是否有系统依赖缺失(如 rJava 需要 Java)\n")
cat("3. 可以尝试单独安装失败的包并查看详细错误信息\n")
}
cat("\n========================================\n")
修改系统文件
修改/home/用户名/shiny-server/omicscloud/etc/config/conf配置文件:
- 用户名都替换一下
- server_href http://localhost/omicscloud/
- dep_content是各app的名称和权限设置:文件前缀名:显示名:类型:权限数字。文件前缀名就是etc/dep里的各个*_func_lib.r *_output.r这种,类型有app\link\module,对应不同的加载方式,module可以参考user management,link可以参考omicsgraph
- db_host localhost ;db_host_string localhost
- ldap_server_url ldap://localhost
- server_data_path /home/lyx/mnt/omicscloud/data/